MicroRNAs as biomarkers in leukemia
Introduction
MicroRNAs (miRNAs) are a novel class of endogenous non-coding RNAs ranging from 19 to 25 nucleotides in size, which regulate gene expression primarily at the post-transcriptional level and thus are involved in many cellular processes, such as cell proliferation, differentiation, and apoptosis (1-3). Recent studies have shown that dysregulated expression of specific miRNAs that function as tumor suppressors or oncogenes is associated with the pathogenesis of human cancers and that specific miRNA expression signatures can be used to effectively classify human tumors (4-6). MiRNAs associated with specific cytogenetic changes and clinical outcomes of different subtypes of leukemia have been reported, demonstrating miRNAs have the potential to be used for clinical diagnosis, prognosis and cancer therapy (7-13).
Human leukemia is classified into acute leukemia and chronic leukemia clinically and pathologically, however, evidence suggests that all leukemia derives from a common leukemic stem cell (LSC) (14). Leukemia is characterized by the abnormal proliferation of blood precursor cells of myeloid or lymphoid origin and the number one cancer killer of children less than 14 years of age (15). Recently, biomarkers of different leukemia subtypes based on genetic, phenotypic, or molecular characteristics have been reported. For examples, the expression of CD38 and ZAP-70 can be used as prognostic marker in chronic lymphocytic leukemia (CLL), while the mutational status of NPM1, FLT3, CEBPA, and MLL are associated with the outcome of treatment for patients with cytogenetically normal acute myeloid leukemia (AML) and can be used to identify AML patients who will benefit from allogeneic stem cell transplantation (SCT) (16,17).
More recently, studies found that miRNAs and long non-coding RNAs (lncRNAs) can be used as biomarkers for diagnosis and prognosis in cancer, revealing that they are a novel source of biomarkers that may be applied to clinical setting (18-21). Here we will review the role of miRNAs in leukemia, mainly focusing on recent findings that suggest the potential of miRNAs as biomarkers for diagnosis and prognosis (Table 1). Moreover, the circulating miRNAs were also emphasized for the fact that they can be detected in body fluids making them suitable to serve as biomarkers.
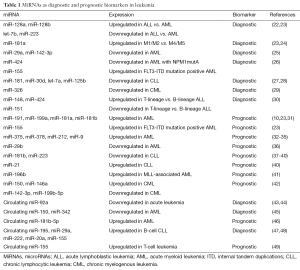
Full table
miRNAs as diagnostic biomarkers
Some studies have revealed that miRNAs may serve as biomarkers in clinical diagnosis. Mi et al. demonstrated that miR-128a, miR-128b, let-7b and miR-223 were the most significant and discriminatory between AML and acute lymphoblastic leukemia (ALL) (22). Moreover, large-scale real-time PCR analysis using two of these four miRNAs in the diagnosis of 98 ALL and AML cases could result in an accuracy rate as high as 97-100%, which provides new potential markers for the classification and diagnosis of ALL and AML (22). Zhu et al. further confirmed that miR-128, let-7b, miR-223 and miR-181a have a diagnosis value in acute leukemia patients (23). Another study demonstrated that a better diagnostic outcome was achieved by combining miR-29a and miR-142-3p with about 90% sensitivity, 100% specificity, indicating that the expression level of miR-142-3p and miR-29a in peripheral blood mononuclear cells (PBMNC) could be used as novel diagnostic marker in AML (25). Debernardi et al. revealed that miR-181a correlated strongly with the AML morphological subtype (24). It was also found that miR-424 was down-regulated in AML patients with NPM1mutA regardless of FLT3 status, while miR-155 showed up-regulation in patients with FLT3 internal tandem duplications (ITD) with or without NPM1 mutations, suggesting that miR-424 and miR-155 deregulation is involved in the pathogenesis of cytogenetically normal AML with NPM1 and FLT3-ITD mutations, respectively (26). Interestingly, another study reported that the expression signatures of miR-22 and its host gene C17orf91 are associated with specific myeloid leukemia subtypes, implicating their potential application for the classification of leukemia (50).
In CLL, Marton et al. identified several miRNAs including miR-181, miR-30d and let-7a that are differentially expressed between CLL lymphocytes and CD19+ control cells (27). Tili et al. found that both aggressive and indolent CLL patients show reduced expression of miR-125b and defined a miR-125b-dependent CLL metabolism-related transcript signature, suggesting miR-125b acts as a master regulator and could be an indicator for the adaptation of cell metabolism to a transformed state in CLL (28). In chronic myelogenous leukemia (CML), downregulation of miR-326 may be a possible mechanism for unrestricted activation of Smo signal transducer of the oncogenic Hh pathway and resulted in elevated cell proliferation and decreased rate of apoptosis in CML CD34(+) cells (29). In ALL, miR-148, miR-151 and miR-424 were identified as discriminative of T-lineage versus B-lineage ALL (30).
MiRNAs as prognostic biomarkers
Accumulating evidence has shown that miRNAs can be used as biomarkers to predict the prognosis of patients. In AML, Garzon et al. demonstrated that a subset of deregulated miRNAs is associated with outcome and AML patients with high expression of miR-191 and miR-199a had significantly worse overall and event free survival than patients with low expression (10). Another study identified a miRNA signature which was associated with event-free survival in cytogenetically normal AML patients with high-risk molecular features (FLT3-ITD, a wild-type NPM1, or both) and found increased miR-181a and miR-181b were significantly associated with a favorable prognosis (31). Zhu et al. also found high expression of miR-181a suggested a better prognosis, while miR-155 can lead to increased myeloid progenitor cells, which may be related to poor prognosis in AML (23). High miR-375 expression was found significantly associated with shorter relapse-free survival (RFS) and overall survival (OS) in pediatric AML patients, demonstrating it may be a novel biomarker to improve the management of pediatric AML patients (32). Overexpression of miR-378 is frequent and may affect treatment outcomes in AML patients, suggesting that miR-378 might have an adverse impact on prognosis in AML (33). Using a large AML patient cohort, miR-212 was found significantly associated with a prolonged OS and RFS, while miR-9 and miR29b indicate poor prognosis in AML (34-36). Using a 45-miRNA profile, Templin et al. can predict the class membership of acute leukemia samples with unknown irradiation status, with accuracies of 100%, supporting that the miRNA expression signature can be used as biomarkers of radiation exposure (51).
It has been found that during the course of CLL, miR-181b decreased in samples of patients with a progressive but not in samples of patients with a stable disease over time, and miR-181b can be as an independent biomarker for the progression of this disease from indolent to aggressive (37,38). Multivariate analysis revealed that the absence of miR-223 was the only independent factor capable of predicting shorter PFS and is a useful biomarker for patients with CLL (39). Another study reported that miR-21 expression levels were significantly higher in CLL patients with poor prognosis and predicted OS, and miR-181b expression levels significantly predicted treatment-free survival, indicating they are prognostic factors (40).
Interestingly, studies showed that a 14-miRNA signature was epigenetically regulated in ALL patients and the methylation status of these miRNAs is a good predictor of ALL outcome after SCT, suggesting that the epigenetic biomarkers have prognostic significance and could improve the stratification of patients into risk groups (52). However, miR-196b resulted in more aggressive leukemic phenotypes and caused much faster leukemogenesis in secondary transplantation than MLL fusion alone in MLL-rearranged leukemia (41). Flamant et al. found increased expression of miR-150 and miR-146a, and reduced expression of miR-142-3p and miR-199b-5p after two weeks of imatinib therapy for CML, suggesting that these miRNAs may serve as a novel clinically useful biomarker in CML (42). Analysis of miR-16 expression level in 72 patients with T-lymphoblastic lymphoma/leukemia showed miR-16 is associated with clinical outcome and may be considered as a potential prognostic marker for T-lymphoblastic lymphoma/leukemia (53).
Circulating miRNAs as diagnostic and prognostic biomarkers
Recent studies have suggested improvements in detecting malignancies using specific extracellular miRNAs in body fluids and circulating miRNAs might be a new class of effective biomarkers (54).
It has been revealed that miR-92a is dramatically decreased in the plasma of acute leukemia patients and the ratio of miR-92a/miR-638 in plasma is very useful for distinguishing leukemia patients from healthy subjects, thus the ratio of miR-92a/miR-638 has strong potential for clinical application as a novel biomarker for detection of leukemia (43). Another study compared miR-92a expression in plasma with its expression in acute leukemia cells and found that the cell to plasma ratio of miR-92a expression was significantly higher in both AML and ALL cells compared with PBMNC from healthy volunteers, implying the ratio of miR-92a expression is a novel blood based leukemia biomarker (44). By analyzing the profile of circulating miRNAs in the plasma of AML patients compared with healthy individuals, it was found that the combination of plasma miR-150 and miR-342 is a novel candidate diagnostic biomarker of AML and potential predictor of relapse (45). Zhi et al. revealed the remarkable ability of a 6-miRNA profile to differentiate between AML and normal controls. Especially, miR-181b-5p level in serum is significantly associated with OS, demonstrating miR-181b-5p may serve as a predictor for OS in AML patients (46).
A recent study reported that circulating miR-195, miR-29a and miR-222 levels are the best classifiers to separate B-cell CLL patients from healthy controls. When combining the levels of several miRNAs, including miR-29a, miR-483–5p, miR-195, miR-185, miR-135a* and miR-15a, it provides good separation between ZAP-70+ and ZAP-70− samples and miR-29a is the single best predictor for ZAP-70 expression status. Notably, the level of circulating miR-20a was determined to correlate reliably with diagnosis-to-treatment time (47). To examine the prognostic role of miR-155, Ferrajoli et al. measured its expression level in plasma samples of 228 newly diagnosed B-cell CLL patients and found plasma level of miR-155 can be used as biomarker to identify patients with CLL that may not respond well to therapy (48). In adult T-cell leukemia, the elevation of plasma miR-155 and the reduction in miR-126 correlate with poor prognosis, indicating they are novel biomarkers for the assessment of disease stage (49).
Conclusions and perspectives
Improving early cancer detection and diagnosis are critical to successful cancer treatment. Studies have revealed miRNAs function as important regulators in gene regulatory networks and exert crucial roles in cancer progression. Mounting evidence showed that miRNAs have great potential as biomarkers and targets for novel therapeutic approaches in the future. More importantly, recent studies demonstrated that circulating miRNA profiles in plasma and serum reflect physiological or pathological conditions, exhibiting they might be a new class of effective biomarkers. So far, unique miRNA patterns and molecular mechanisms of some miRNAs in different cancers and diseases have been fully studied. It should be noted that some miRNAs have similar expression levels and functions in different cancers or cancer subtypes. It is necessary to further validate the reproducibility and reliability of miRNAs as biomarkers for diagnosis and prognosis in a large scale of clinical samples, making it possible to widely use these miRNAs as biomarkers to detect multiple cancers. It remains a significant challenge to translate the basic research to the clinical setting, however, the important roles of miRNAs in cancer have shed light on their potential applications and will ultimately provide novel strategies for cancer diagnosis and therapy.
Acknowledgements
Funding: This work was supported by National Natural Science Foundation of China (No. 81071638, No. 81300398, No. 31170676) and the funds from Scientific Research Foundation for Returned Overseas Scholars of Guangdong Medical College, China (XB1377, XB1378).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281-97. [PubMed]
- Ambros V. The functions of animal microRNAs. Nature 2004;431:350-5. [PubMed]
- Lim LP, Lau NC, Garrett-Engele P, et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005;433:769-73. [PubMed]
- Chen CZ. MicroRNAs as oncogenes and tumor suppressors. N Engl J Med 2005;353:1768-71. [PubMed]
- Esquela-Kerscher A, Slack FJ. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 2006;6:259-69. [PubMed]
- Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature 2005;435:834-8. [PubMed]
- Zhang H, Luo XQ, Zhang P, et al. MicroRNA patterns associated with clinical prognostic parameters and CNS relapse prediction in pediatric acute leukemia. PLoS One 2009;4:e7826. [PubMed]
- Labbaye C, Testa U. The emerging role of MIR-146A in the control of hematopoiesis, immune function and cancer. J Hematol Oncol 2012;5:13. [PubMed]
- Calin GA, Ferracin M, Cimmino A, et al. A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 2005;353:1793-801. [PubMed]
- Garzon R, Volinia S, Liu CG, et al. MicroRNA signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood 2008;111:3183-9. [PubMed]
- Gimenes-Teixeira HL, Lucena-Araujo AR, Dos Santos GA, et al. Increased expression of miR-221 is associated with shorter overall survival in T-cell acute lymphoid leukemia. Exp Hematol Oncol 2013;2:10. [PubMed]
- Jongen-Lavrencic M, Sun SM, Dijkstra MK, et al. MicroRNA expression profiling in relation to the genetic heterogeneity of acute myeloid leukemia. Blood 2008;111:5078-85. [PubMed]
- Fernando TR, Rodriguez-Malave NI, Rao DS. MicroRNAs in B cell development and malignancy. J Hematol Oncol 2012;5:7. [PubMed]
- Gilliland DG, Jordan CT, Felix CA. The molecular basis of leukemia. Hematology Am Soc Hematol Educ Program 2004.80-97. [PubMed]
- Lightfoot T. Aetiology of childhood leukemia. Bioelectromagnetics 2005.Suppl 7:S5-11. [PubMed]
- Chiorazzi N. Implications of new prognostic markers in chronic lymphocytic leukemia. Hematology Am Soc Hematol Educ Program 2012;2012:76-87.
- Schlenk RF, Döhner K, Krauter J, et al. Mutations and treatment outcome in cytogenetically normal acute myeloid leukemia. N Engl J Med 2008;358:1909-18. [PubMed]
- Zhang H, Chen Z, Wang X, et al. Long non-coding RNA: a new player in cancer. J Hematol Oncol 2013;6:37. [PubMed]
- Van Roosbroeck K, Pollet J, Calin GA. miRNAs and long noncoding RNAs as biomarkers in human diseases. Expert Rev Mol Diagn 2013;13:183-204. [PubMed]
- Sun YM, Lin KY, Chen YQ. Diverse functions of miR-125 family in different cell contexts. J Hematol Oncol 2013;6:6. [PubMed]
- Hassan O, Ahmad A, Sethi S, et al. Recent updates on the role of microRNAs in prostate cancer. J Hematol Oncol 2012;5:9. [PubMed]
- Mi S, Lu J, Sun M, et al. MicroRNA expression signatures accurately discriminate acute lymphoblastic leukemia from acute myeloid leukemia. Proc Natl Acad Sci U S A 2007;104:19971-6. [PubMed]
- Zhu YD, Wang L, Sun C, et al. Distinctive microRNA signature is associated with the diagnosis and prognosis of acute leukemia. Med Oncol 2012;29:2323-31. [PubMed]
- Debernardi S, Skoulakis S, Molloy G, et al. MicroRNA miR-181a correlates with morphological sub-class of acute myeloid leukaemia and the expression of its target genes in global genome-wide analysis. Leukemia 2007;21:912-6. [PubMed]
- Wang F, Wang XS, Yang GH, et al. miR-29a and miR-142-3p downregulation and diagnostic implication in human acute myeloid leukemia. Mol Biol Rep 2012;39:2713-22. [PubMed]
- Faraoni I, Laterza S, Ardiri D, et al. MiR-424 and miR-155 deregulated expression in cytogenetically normal acute myeloid leukaemia: correlation with NPM1 and FLT3 mutation status. J Hematol Oncol 2012;5:26. [PubMed]
- Marton S, Garcia MR, Robello C, et al. Small RNAs analysis in CLL reveals a deregulation of miRNA expression and novel miRNA candidates of putative relevance in CLL pathogenesis. Leukemia 2008;22:330-8. [PubMed]
- Tili E, Michaille JJ, Luo Z, et al. The down-regulation of miR-125b in chronic lymphocytic leukemias leads to metabolic adaptation of cells to a transformed state. Blood 2012;120:2631-8. [PubMed]
- Babashah S, Sadeghizadeh M, Hajifathali A, et al. Targeting of the signal transducer Smo links microRNA-326 to the oncogenic Hedgehog pathway in CD34+ CML stem/progenitor cells. Int J Cancer 2013;133:579-89. [PubMed]
- Fulci V, Colombo T, Chiaretti S, et al. Characterization of B- and T-lineage acute lymphoblastic leukemia by integrated analysis of MicroRNA and mRNA expression profiles. Genes Chromosomes Cancer 2009;48:1069-82. [PubMed]
- Marcucci G, Radmacher MD, Maharry K, et al. MicroRNA expression in cytogenetically normal acute myeloid leukemia. N Engl J Med 2008;358:1919-28.
- Wang Z, Hong Z, Gao F, et al. Upregulation of microRNA-375 is associated with poor prognosis in pediatric acute myeloid leukemia. Mol Cell Biochem 2013;383:59-65. [PubMed]
- Qian J, Lin J, Qian W, et al. Overexpression of miR-378 is frequent and may affect treatment outcomes in patients with acute myeloid leukemia. Leuk Res 2013;37:765-8. [PubMed]
- Sun SM, Rockova V, Bullinger L, et al. The prognostic relevance of miR-212 expression with survival in cytogenetically and molecularly heterogeneous AML. Leukemia 2013;27:100-6. [PubMed]
- Maki K, Yamagata T, Sugita F, et al. Aberrant expression of MIR9 indicates poor prognosis in acute myeloid leukaemia. Br J Haematol 2012;158:283-5. [PubMed]
- Xiong Y, Li Z, Ji M, et al. MIR29B regulates expression of MLLT11 (AF1Q), an MLL fusion partner, and low MIR29B expression associates with adverse cytogenetics and poor overall survival in AML. Br J Haematol 2011;153:753-7. [PubMed]
- Visone R, Veronese A, Rassenti LZ, et al. miR-181b is a biomarker of disease progression in chronic lymphocytic leukemia. Blood 2011;118:3072-9. [PubMed]
- Visone R, Veronese A, Balatti V, et al. MiR-181b: new perspective to evaluate disease progression in chronic lymphocytic leukemia. Oncotarget 2012;3:195-202. [PubMed]
- Zhou K, Yi S, Yu Z, et al. MicroRNA-223 expression is uniformly down-regulated in B cell lymphoproliferative disorders and is associated with poor survival in patients with chronic lymphocytic leukemia. Leuk Lymphoma 2012;53:1155-61. [PubMed]
- Rossi S, Shimizu M, Barbarotto E, et al. microRNA fingerprinting of CLL patients with chromosome 17p deletion identify a miR-21 score that stratifies early survival. Blood 2010;116:945-52. [PubMed]
- Li Z, Huang H, Chen P, et al. miR-196b directly targets both HOXA9/MEIS1 oncogenes and FAS tumour suppressor in MLL-rearranged leukaemia. Nat Commun 2012;3:688. [PubMed]
- Flamant S, Ritchie W, Guilhot J, et al. Micro-RNA response to imatinib mesylate in patients with chronic myeloid leukemia. Haematologica 2010;95:1325-33. [PubMed]
- Tanaka M, Oikawa K, Takanashi M, et al. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS One 2009;4:e5532. [PubMed]
- Ohyashiki JH, Umezu T, Kobayashi C, et al. Impact on cell to plasma ratio of miR-92a in patients with acute leukemia: in vivo assessment of cell to plasma ratio of miR-92a. BMC Res Notes 2010;3:347. [PubMed]
- Fayyad-Kazan H, Bitar N, Najar M, et al. Circulating miR-150 and miR-342 in plasma are novel potential biomarkers for acute myeloid leukemia. J Transl Med 2013;11:31. [PubMed]
- Zhi F, Cao X, Xie X, et al. Identification of circulating microRNAs as potential biomarkers for detecting acute myeloid leukemia. PLoS One 2013;8:e56718. [PubMed]
- Moussay E, Wang K, Cho JH, et al. MicroRNA as biomarkers and regulators in B-cell chronic lymphocytic leukemia. Proc Natl Acad Sci U S A 2011;108:6573-8. [PubMed]
- Ferrajoli A, Shanafelt TD, Ivan C, et al. Prognostic value of miR-155 in individuals with monoclonal B-cell lymphocytosis and patients with B chronic lymphocytic leukemia. Blood 2013;122:1891-9. [PubMed]
- Ishihara K, Sasaki D, Tsuruda K, et al. Impact of miR-155 and miR-126 as novel biomarkers on the assessment of disease progression and prognosis in adult T-cell leukemia. Cancer Epidemiol 2012;36:560-5. [PubMed]
- Wang J, Xiang G, Zhang K, et al. Expression signatures of intragenic miRNAs and their corresponding host genes in myeloid leukemia cells. Biotechnol Lett 2012;34:2007-15. [PubMed]
- Templin T, Paul S, Amundson SA, et al. Radiation-induced micro-RNA expression changes in peripheral blood cells of radiotherapy patients. Int J Radiat Oncol Biol Phys 2011;80:549-57. [PubMed]
- Martín-Palanco V, Rodríguez G, Martín C, et al. microRNA methylation profile has prognosis impact in acute lymphoblastic leukemia patients undergoing stem cell transplantation. Biol Blood Marrow Transplant 2011;17:745-8. [PubMed]
- Xi Y, Li J, Zan L, et al. Micro-RNA-16 expression in paraffin-embedded specimen correlates with overall survival of T-lymphoblastic lymphoma/leukemia. Hum Pathol 2013;44:1011-6. [PubMed]
- Madhavan D, Cuk K, Burwinkel B, et al. Cancer diagnosis and prognosis decoded by blood-based circulating microRNA signatures. Front Genet 2013;4:116. [PubMed]
Cite this article as: Wang X, Zhu B, Huang Z, Chen L, He Z, Zhang H. MicroRNAs as biomarkers in leukemia. Stem Cell Investig 2014;1:11.


